Cell Ranger ARC2.0, printed on 03/03/2025
The Illumina NovaSeq control software v1.8 upgrade affects cellranger-arc mkfastq's ability to autodetect the i5 (Index 2) orientation due to Illumina's reagent name changes in the recipe XML file. This results in a significant number of GEX reads going into Undetermined and there will be no error. ATAC reads are unaffected. The issue has been fixed in Cell Ranger ARC v2.0.2. If you are using an older version of Cell Ranger ARC, several solutions are provided in this Knowledge Base article.
|
The cellranger-arc workflow starts by demultiplexing the Illumina sequencer's base call files (BCLs) for each flow cell directory (ATAC or Gene Expression) into FASTQ files. 10x Genomics has developed cellranger-arc mkfastq, a pipeline that wraps Illumina's bcl2fastq and provides a number of convenient features in addition to the features of bcl2fastq:
--use-bases-mask.The Multiome ATAC library is single-indexed while the Multiome GEX library is
dual-indexed. cellranger-arc mkfastq can auto-detect the
type of flow cell based on the length of the i5 index read and selects the
appropriate mode depending on the sample indexes used, and enables index-hopping
filtering automatically for dual-indexed flow cells. For example, a Multiome GEX
library prepared with the Dual Index Kit TT Set A, well A1 can be specified in
the sample sheet as "SI-TT-A1", and cellranger-arc mkfastq
will recognize the i7 and i5 indices as GTAACATGCG and AGTGTTACCT,
respectively. Similarly for a Multiome ATAC library prepared with Single Index
Kit N Set A, well A1 can be specified in the sample sheet as "SI-NA-A1", and
cellranger-arc mkfastq will recognize the four i7 indexes
(AAACGGCG, CCTACCAT, GGCGTTTC, and TTGTAAGA) and merge the resulting FASTQ
files.
The compute workflow begins with running one instance of cellranger-arc mkfastq for each flow cell of data being analyzed. The same command cellranger-arc mkfastq can be used to demultiplex ATAC and GEX flow cells. Once the ATAC flow cell(s) and GEX flow cell(s) are successfully demultiplexed, one instance of cellranger-arc count is run for each paired Multiome ATAC and GEX library; independent of the number of sequencing runs of each library. Specific examples are described below.
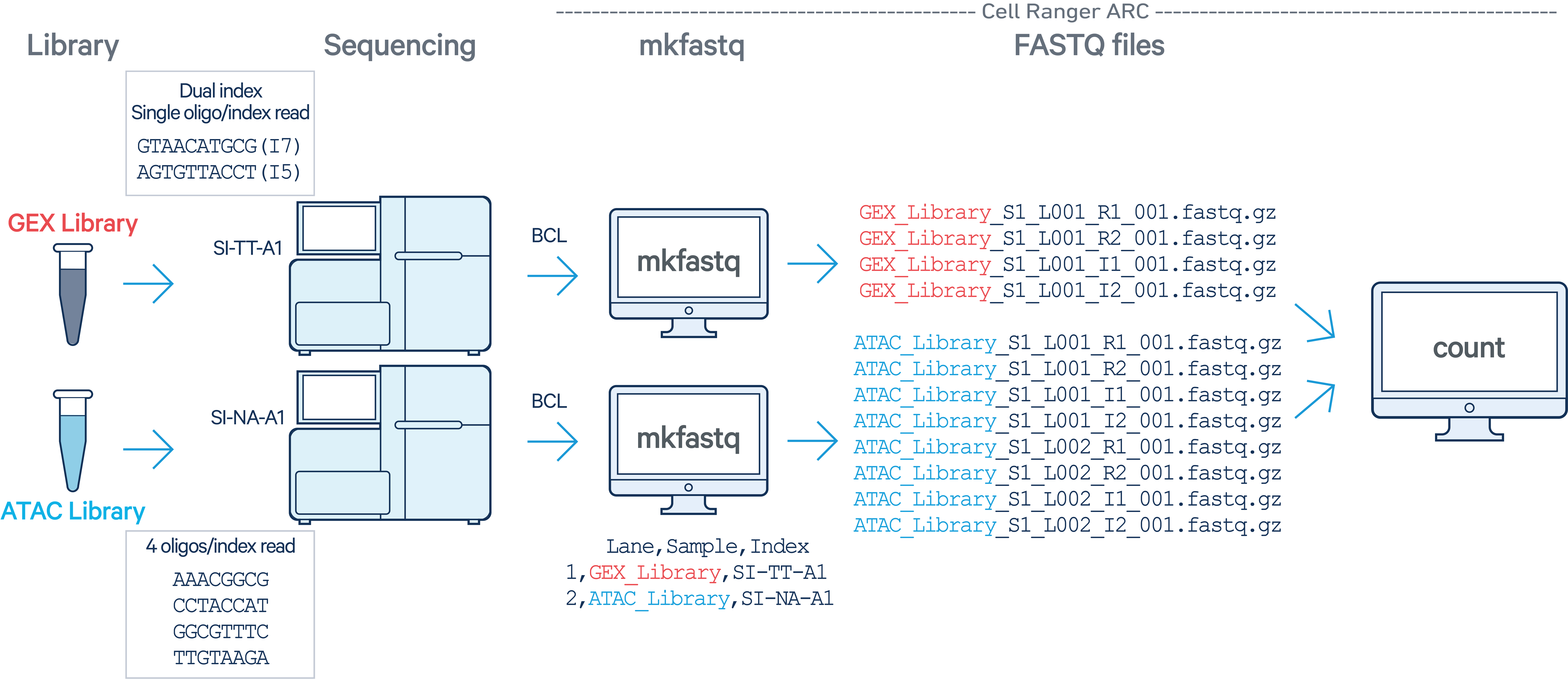
In this example, a Multiome ATAC library (with sample index SI-NA-A1) and a Multiome GEX library (with sample index SI-TT-A1) were processed on different flow cells. The GEX library was processed on lane 1 of its flow cell, whereas the ATAC library was processed on lane 2 of its flow cell. Then, a separate instance of cellranger-arc mkfastq is run for each library, and all resultant FASTQ files are processed though a single instance of cellranger-arc count.
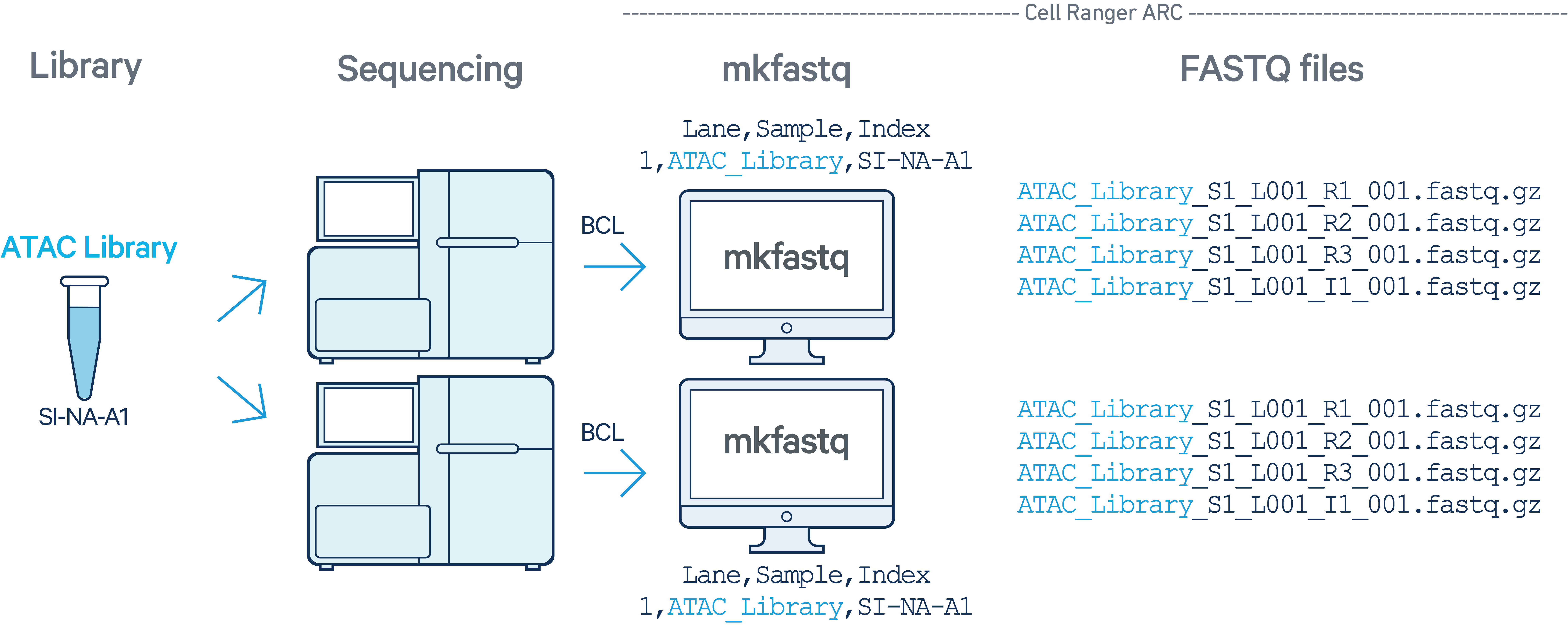
In this example, one Multiome ATAC library with sample index SI-NA-A1 was sequenced on two flow cells. The cellranger-arc count pipeline cannot process ATAC libraries alone.
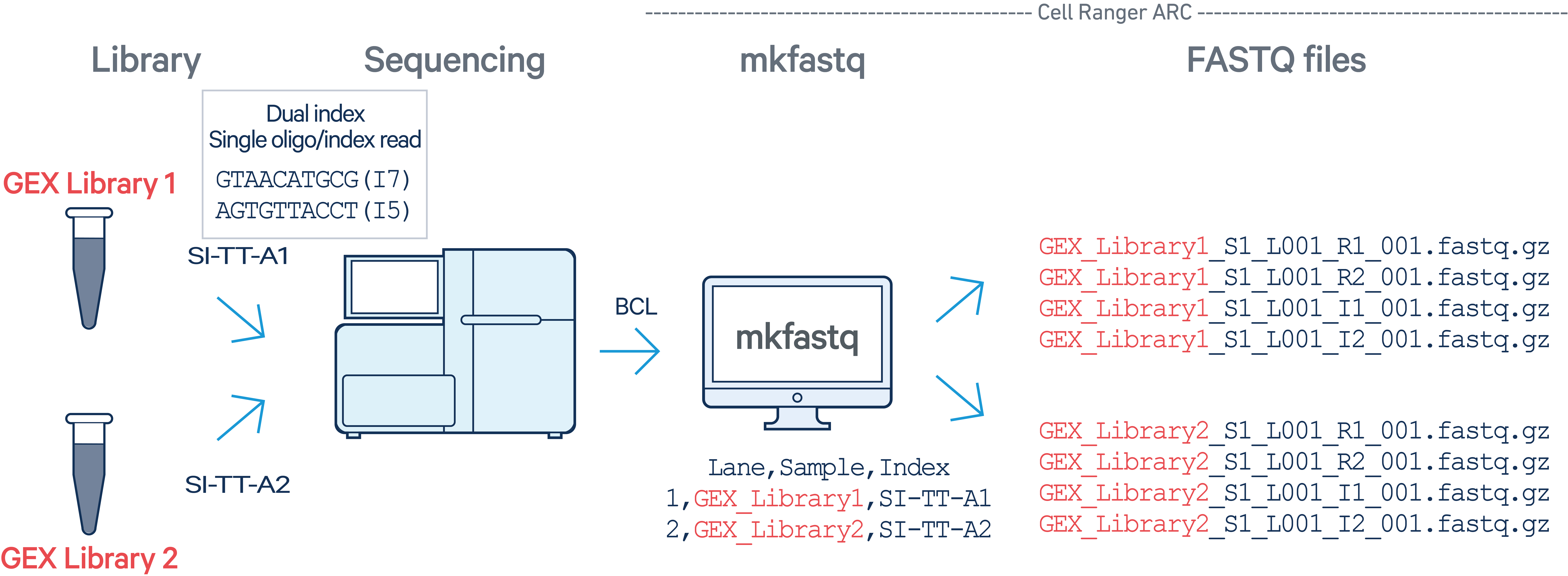
In this example, two Multiome GEX libraries (each processed through a separate GEM well with sample indices SI-TT-A1 and SI-TT-A2) are multiplexed on a single flow cell. GEX Library 1 was processed on lane 1 and GEX Library 2 was processed on lane 2 of the same flow cell. The cellranger-arc count pipeline cannot process GEX libraries alone.
The cellranger-arc mkfastq pipeline accepts additional options beyond
those shown in the table below because it is a wrapper around
bcl2fastq. Consult the User Guide for Illumina's
bcl2fastq
for more information.
| Parameter | Function |
|---|---|
--run | Required. The path of Illumina BCL run folder. |
--id | Optional; defaults to the name of the flow cell referred to by --run. Name of the folder created by mkfastq. |
--samplesheet | Optional. Path to an Illumina Experiment Manager-compatible sample sheet which contains 10x Genomics sample index names (e.g., SI-NA-A1 or SI-TT-A12) in the sample index column. All other information, such as sample names and lanes, should be in the sample sheet. |
--sample-sheet | Optional. Equivalent to --samplesheet above. |
--csv | Optional. Path to a simple CSV with lane, sample, and index columns, which describe the way to demultiplex the flow cell. The index column should contain a 10x Genomics sample dual-index name (e.g., SI-TT-A12). This is an alternative to the Illumina IEM samplesheet, and will be ignored if --samplesheet is specified. |
--simple-csv | |
--lanes | bcl2fastq option. Comma-delimited series of lanes to demultiplex (e.g. 1,3). Use this if you have a sample sheet for an entire flow cell but only want to generate a few lanes for further 10x Genomics analysis. |
--use-bases-mask | bcl2fastq option. Same meaning as for bcl2fastq. Use to clip extra bases off a read if you ran extra cycles for QC. |
--delete-undetermined | bcl2fastq option. Delete the Undetermined FASTQs generated by bcl2fastq. Useful if you are demultiplexing a small number of samples from a large flow cell. |
--barcode-mismatches | bcl2fastq option. Same meaning as for bcl2fastq. Use this option to change the number of allowed mismatches per index adapter (0, 1, 2). Default: 1. |
--output-dir | bcl2fastq option. Generate FASTQ output in a path of your own choosing, instead of flow_cell_id/outs/fastq_path. |
--project | bcl2fastq option. Custom project name, to override the sample sheet or to use in conjunction with the --csv argument. |
--jobmode | Martian option. Job manager to use. Valid options: local (default), sge, lsf, slurm or a .template file. |
--localcores | Martian option. Set max cores the pipeline may request at one time. Only applies when --jobmode=local. |
--localmem | Martian option. Set max GB the pipeline may request at one time. Only applies when --jobmode=local. |
The cellranger-arc mkfastq pipeline recognizes two file formats for
describing samples: a simple, three-column CSV format, or the Illumina
Experiment Manager (IEM) sample sheet format used by bcl2fastq. Both these formats are illustrated with a Multiome ATAC flow cell and Multiome GEX flow cell example.
The example (tiny-bcl-atac) dataset is solely designed to demo the cellranger-arc mkfastq pipeline. It cannot be used to run downstream pipelines (e.g. cellranger-arc count).
To follow along, do the following:
cellranger-arc-tiny-bcl-atac-1.0.0.tar.gz and
cellranger-arc-tiny-bcl-gex-1.0.0.tar.gz tar files in a convenient location.A simple CSV sample sheet is recommended for most sequencing experiments.
The simple CSV format has only three columns (Lane, Sample, Index), and is thus
less prone to formatting errors. You can see an example of this in
cellranger-arc-tiny-bcl-atac-simple-1.0.0.csv:
Lane,Sample,Index 1,test_sample_atac,SI-NA-A1
and in cellranger-arc-tiny-bcl-gex-simple-1.0.0.csv:
Lane,Sample,Index 1,test_sample_gex,SI-TT-A1
Here are the options for each column:
| Lane | Which lane(s) of the flow cell to process. Can be either a single lane, a range (e.g., 2-4) or '*' for all lanes in the flow cell. |
| Sample | The name of the sample. This name is the prefix to all the generated FASTQs, and corresponds to the --sample argument in all downstream 10x Genomics pipelines.Sample names must conform to the Illumina bcl2fastq naming requirements. Only letters, numbers, underscores, and hyphens are allowed; no other symbols, including dots ("."), are allowed. |
| Index | The 10x Genomics sample index that was used in library construction, e.g., SI-TT-A1 for a Dual-Indexed Multiome GEX library, or SI-NA-A1 for a Multiome ATAC library. |
To run cellranger-arc mkfastq with a simple layout CSV, use the --csv
argument. Here's how to run cellranger-arc mkfastq on the
tiny-bcl-atac sequencing run with the simple layout (replace code in red with the path to cellranger-arc-tiny-bcl-atac on your system):
$ cellranger-arc mkfastq --id=tiny-bcl-atac \ --run=/path/to/cellranger-arc-tiny-bcl-atac-1.0.0 \ --csv=cellranger-arc-tiny-bcl-atac-simple-1.0.0.csv cellranger-arc mkfastq (2.0.2) Copyright (c) 2020 10x Genomics, Inc. All rights reserved. ------------------------------------------------------------------------------- Martian Runtime - v4.0.5 Running preflight checks (please wait)... yyyy-mm-dd hh:mm:ss [runtime] (ready) ID.tiny-bcl-atac.MAKE_FASTQS_CS.MAKE_FASTQS.PREPARE_SAMPLESHEET yyyy-mm-dd hh:mm:ss [runtime] (split_complete) ID.tiny-bcl-atac.MAKE_FASTQS_CS.MAKE_FASTQS.PREPARE_SAMPLESHEET yyyy-mm-dd hh:mm:ss [runtime] (run:local) ID.tiny-bcl-atac.MAKE_FASTQS_CS.MAKE_FASTQS.PREPARE_SAMPLESHEET.fork0.chnk0.main yyyy-mm-dd hh:mm:ss [runtime] (chunks_complete) ID.tiny-bcl-atac.MAKE_FASTQS_CS.MAKE_FASTQS.PREPARE_SAMPLESHEET ...
The cellranger-arc mkfastq pipeline can also be run with a
sample sheet in the Illumina Experiment Manager (IEM) format. An IEM sample sheet has several fields specific to running on Illumina platforms, including a [Data] section where sample and index information is specified. cellranger-arc mkfastq supports listing either index set names or the oligo sequences.
|
Do not trim adapters during demultiplexing. Leave these settings blank. Trimming adapters from reads can potentially damage the 10x barcodes and the UMIs, resulting in pipeline failure or data loss.
If you are using an Illumina sample sheet for demultiplexing with bcl2fastq, BCL Convert or our mkfastq pipeline, please remove these lines under the [Settings]section: Adapter or AdapterRead1 or AdapterRead2.
|
Example [Data] section for a dual-indexed Multiome GEX flow cell
Version 1: "SI-TT-A1" refers to a 10x Genomics dual-indexed library sample index, so mkfastq auto-detects that this is a dual-index sample. In this example, only reads from lane 1 will be used. To demultiplex the given sample index across all lanes, omit the lanes column entirely.
[Data] Lane,Sample_ID,index 1,test_sample,SI-TT-A1
Version 2: The index sequences for "SI-TT-A1" are specified in the two index and index2 columns.
[Data] Lane,Sample_ID,index,index2 1,test_sample,GTAACATGCG,AGGTAACACT
Example [Data] section for a single-indexed Multiome ATAC flow cell
Version 1: "SI-NA-A1" refers to a 10x Genomics single-indexed sample index consisting of a set of four oligo sequences.
[Data] Sample_ID,index test_sample_miseq,SI-NA-A1
Version 2: The four index sequences for "SI-NA-A1" are specified in separate rows under the index column.
[Data] Lane,Sample_ID,index 1,sample1,AAACGGCG 1,sample1,CCTACCAT 1,sample1,GGCGTTTC 1,sample1,TTGTAAGA
Sample names must conform to the Illumina bcl2fastq naming requirements. Only letters, numbers, underscores, and hyphens are allowed. No other symbols, including dots ("."), are allowed.
Also note that while an authentic IEM sample sheet will contain other sections
above the [Data] section, these are optional for demultiplexing. To avoid data loss from trimming, we do not recommend including adapter sequences in the [Settings] section of the sample sheet (see this article for details). For demultiplexing an existing run with cellranger-arc
mkfastq, only the [Data] section is required.
Next, run the cellranger-arc mkfastq pipeline, using the
--samplesheet argument (replace code in red with the path to tiny_bcl on your system):
$ cellranger-arc mkfastq --id=tiny-bcl-atac \ --run=/path/to/tiny-bcl-atac \ --samplesheet=cellranger-arc-tiny-bcl-atac-samplesheet-1.0.0.csv cellranger-arc mkfastq (2.0.2) Copyright (c) 2020 10x Genomics, Inc. All rights reserved. ------------------------------------------------------------------------------- Martian Runtime - v4.0.5 Running preflight checks (please wait)... yyyy-mm-dd hh:mm:ss [runtime] (ready) ID.tiny-bcl-atac.MAKE_FASTQS_CS.MAKE_FASTQS.PREPARE_SAMPLESHEET yyyy-mm-dd hh:mm:ss [runtime] (split_complete) ID.tiny-bcl-atac.MAKE_FASTQS_CS.MAKE_FASTQS.PREPARE_SAMPLESHEET yyyy-mm-dd hh:mm:ss [runtime] (run:local) ID.tiny-bcl-atac.MAKE_FASTQS_CS.MAKE_FASTQS.PREPARE_SAMPLESHEET.fork0.chnk0.main yyyy-mm-dd hh:mm:ss [runtime] (chunks_complete) ID.tiny-bcl-atac.MAKE_FASTQS_CS.MAKE_FASTQS.PREPARE_SAMPLESHEET ...
If you encounter any preflight errors, refer to the Troubleshooting page.
Once the cellranger-arc mkfastq pipeline has successfully
completed, the output can be found in a new folder named with the value
provided to cellranger-arc mkfastq in the --id option
(if not specified, defaults to the name of the flow cell):
$ cellranger-arc mkfastq --id=tiny-bcl-atac \ --run=/path/to/tiny-bcl-atac \ --samplesheet=cellranger-arc-tiny-bcl-atac-samplesheet-1.0.0.csv cellranger-arc mkfastq (2.0.2) Copyright (c) 2020 10x Genomics, Inc. All rights reserved. ------------------------------------------------------------------------------- Martian Runtime - v4.0.5 ... Pipestance completed successfully! yyyy-mm-dd hh:mm:ss Shutting down. Saving pipestance info to "tiny-bcl-atac/tiny-bcl-atac.mri.tgz" $ ls -l drwxrwxr-x 4 jdoe jdoe 4096 Aug 29 15:29 tiny-bcl-atac
The key output files can be found in outs/fastq_path, and are
organized in the same manner as a conventional bcl2fastq run:
$ ls -l tiny-bcl-atac/outs/fastq_path/ total 31744 drwxrwxr-x 3 jdoe jdoe 24 Sep 7 22:49 p1 drwxrwxr-x 3 jdoe jdoe 26 Sep 7 22:48 Reports drwxrwxr-x 2 jdoe jdoe 193 Sep 7 22:48 Stats -rw-rw-r-- 1 jdoe jdoe 3806257 Sep 7 22:48 Undetermined_S0_L001_I1_001.fastq.gz -rw-rw-r-- 1 jdoe jdoe 967448 Sep 7 22:48 Undetermined_S0_L001_R1_001.fastq.gz -rw-rw-r-- 1 jdoe jdoe 5773976 Sep 7 22:48 Undetermined_S0_L001_R2_001.fastq.gz -rw-rw-r-- 1 jdoe jdoe 12635207 Sep 7 22:48 Undetermined_S0_L001_R3_001.fastq.gz $ tree tiny-bcl-atac/outs/fastq_path/tiny-bcl-atac/ tiny-bcl-atac/outs/fastq_path/p1 └── s1 ├── test_sample_miseq_S1_L001_I1_001.fastq.gz ├── test_sample_miseq_S1_L001_R1_001.fastq.gz ├── test_sample_miseq_S1_L001_R2_001.fastq.gz └── test_sample_miseq_S1_L001_R3_001.fastq.gz
This example was produced with a sample sheet that included p1 as
the Sample_Project, so the directory containing the sample folders is named
p1. If a Sample_Project wasn't specified, or if a simple layout CSV
file was used (which does not have a Sample_Project column), the directory
containing the sample folders would be named according to the flow cell ID
instead.
To remove the Undetermined FASTQs from the output, you can run mkfastq with the
--delete-undetermined flag. To see all
cellranger-arc mkfastq options, run
cellranger-arc mkfastq --help.
If the pipeline crashes while running cellranger-arc
mkfastq, upload this tarball (with the extension .mri.tgz) found in your output
directory. Replace the code in red with your email
$ cellranger-arc upload [email protected] jobid.mri.tgz
where jobid is what you input into the --id option of
mkfastq (if not specified, defaults to the ID of the flow cell).
This tarball
contains numerous diagnostic logs that we can use for debugging.
You will receive an automated email from 10x Genomics. If not, email [email protected]. For the fastest service, respond with the following:
RunInfo.xml and runParameters.xml files from your BCL directory.