Cell Ranger7.0, printed on 11/05/2024
In this part of the analysis walk-through, we use Loupe V(D)J Browser to look at clonotypes within a gene expression cluster, and compare clonotype distributions between clusters. You need these files to proceed:
If you are new to Loupe V(D)J Browser, you may want to read the Loupe V(D)J Browser tutorial before continuing.
To look at clonotypes within gene expression clusters in Loupe V(D)J Browser, first download one or more V(D)J .vloupe files, and then import the gene expression .cloupe file.
To start, open Loupe V(D)J Browser, and click on Browse for a Loupe V(D)J Browser File.
Choose the LungTumorT dataset and press Load. You see the
following after loading:
Next, click on the folder icon on the top toolbar, and select Load clusters from cloupe file.
Choose the LungTumorGEX file that you either downloaded from the previous
exercise, or from the link above. After importing gene expression data, you see an icon next to the .vloupe file description for which corresponding gene expression data has been loaded.
The icon signals that you can filter clonotypes in that file by cluster.
Select the Cluster option from the Filter By field below the clonotype list. Type 'CD8' in the Enter Cluster Name box. The CD8+ Cytotoxic T Cells cluster appears from the autocomplete list. Select it to limit the clonotype list to cells in that cluster. Clicking on each chain in the list shows you the mutations (vertical lines) against the combined V(D)J reference for the chain.
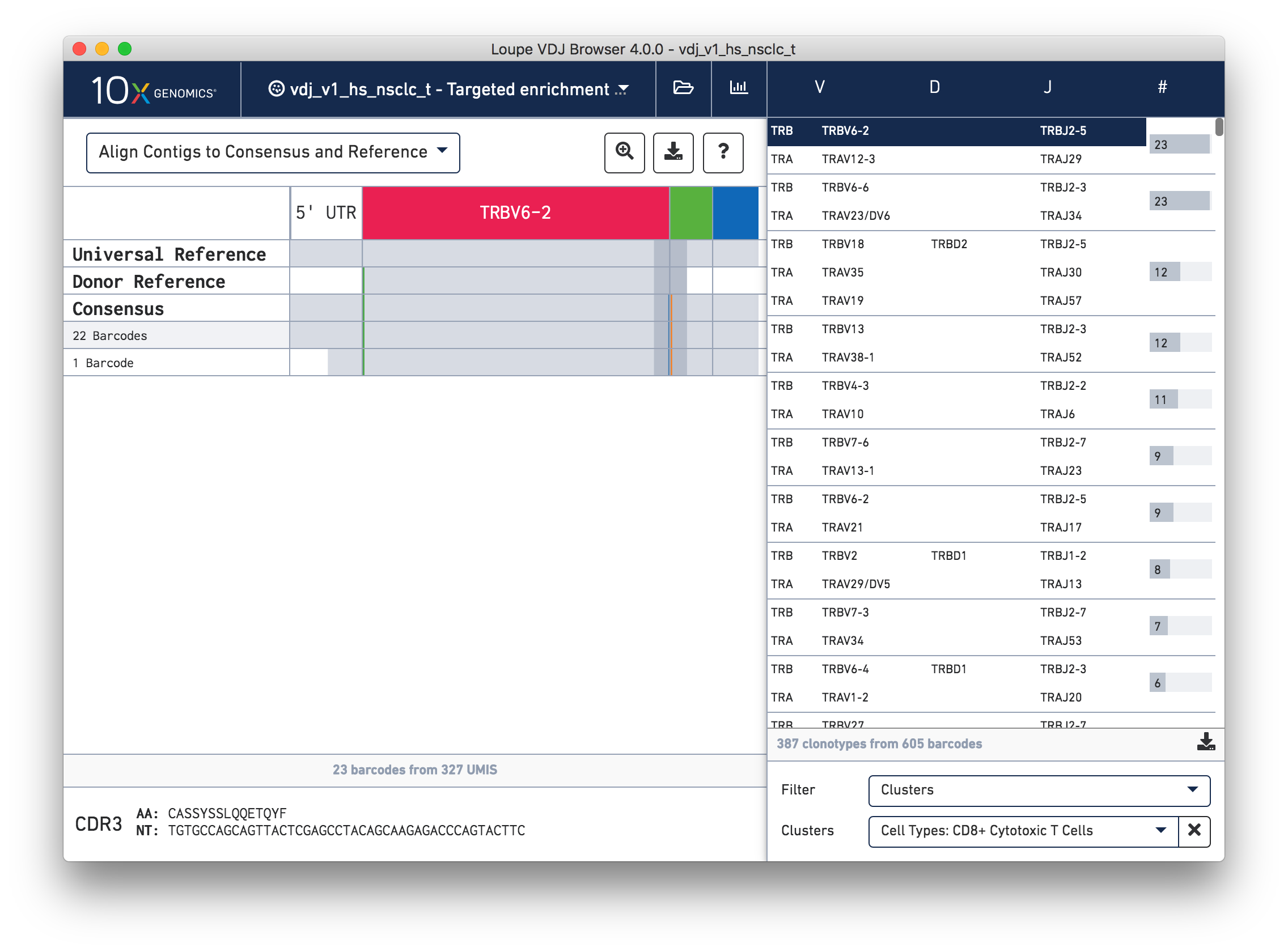
Now that you have seen how to combine V(D)J and Gene Expression data with Loupe Browser and Loupe V(D)J Browser, you may have ideas about how to use the Chromium™ Single Cell V(D)J Solution for your own experiments. If you have any questions or feedback about software or other parts of the integrated V(D)J and Gene Expression workflow, contact [email protected].